New study provides a unique resource for understanding how environmental exposures in early life affect our health
Researchers now have a unique resource for identifying new biomarkers of environmental exposures in early life and understanding their health effects. This is thanks to a study led by the Barcelona Institute for Global Health (ISGlobal), an institution supported by la Caixa Foundation, which systematically documented all associations between a wide range of early life exposures and molecular profiles at different levels, including the epigenome (DNA methylation), transcriptome (gene expression) and metabolome (metabolites).
The findings, which are part of the ATHLETE project, have been published in Nature Communications.
Health depends greatly on the environment. In fact, 70 to 90% of the risk of developing a disease is determined by the exposome, a multitude of environmental factors (i.e., non-genetic factors) to which people are exposed throughout life. And yet, scientists still have limited knowledge on these environmental hazards, how they interact, and what biological processes they trigger.
“Early life is a particularly important period, since exposures during these developmentally vulnerable periods may have pronounced effects at the molecular level, which may not be clinically detectable until adulthood,” explains Martine Vrijheid, Head of the Childhood and Environment Program at ISGlobal.
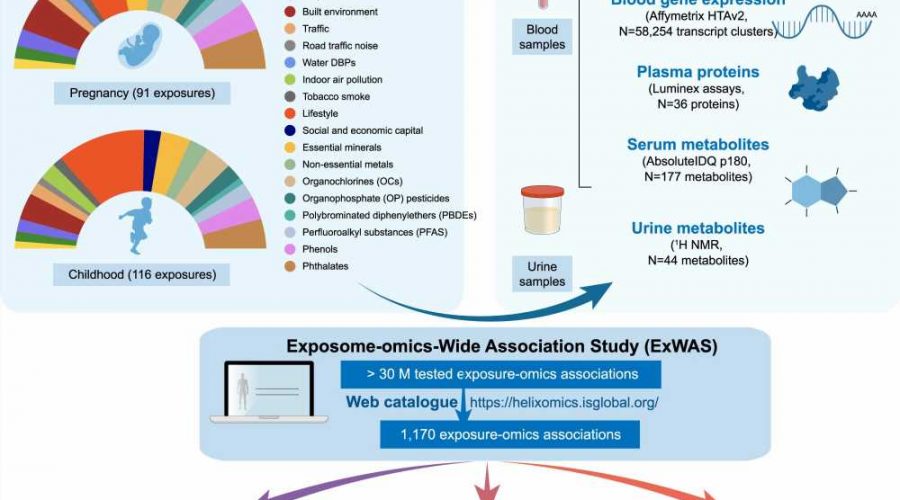
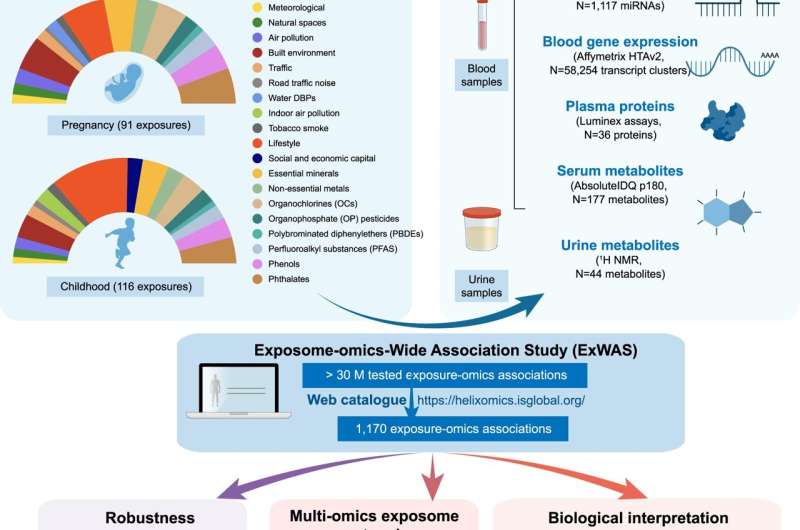
In this study, the research team led by Vrijheid aimed to associate multiple chemical, outdoor, social and lifestyle exposures (92 in pregnancy and 116 when the children were 6 to 11 years old), with molecular profiles in the same children (DNA methylation and gene transcription in blood, plasma proteins, and metabolites in serum and urine). The analysis included 1,301 mother-child pairs of the Human Early Life Exposome (HELIX) project, a long-term cohort study in six European countries (Spain, U.K., France, Lithuania, Norway and Greece).
“The high-performance computing, using massive parallel computers, allowed us to overcome one of the main challenges faced in big ‘omics’ data analyses,” says Juan R Gonzalez, senior co-author. The analysis identified 1,170 significant associations (249 in pregnancy and 921 in childhood) which provide insights into potential biological responses and sources of exposure. Pregnancy exposures, such as maternal smoking, the heavy metal cadmium, or the trace mineral molybdenum, were mostly associated with changes in DNA methylation. In contrast, childhood exposures were associated with signatures at all molecular levels, most particularly with metabolites in serum. The findings revealed, for example, that children are exposed to chemical pollutants through their diet.
“We identified novel multi-omics associations with childhood exposure to essential trace elements, weather conditions, indoor air quality, and phthalates and parabens,” says Léa Maitre, first author. “By visualizing these associations as networks, we can better understand if a given molecular profile is connected to several exposures or vice versa, and thereby identify potential biological pathways,” she adds.
Indeed, the findings provide plausible mechanisms of disease for six groups of exposures: copper, tobacco smoke, indoor air quality during childhood, persistent organic pollutants, phthalates and parabens, and weather conditions. For example, child exposure to copper was associated with almost 90 molecular features, including increased levels of C-reactive protein (an inflammation marker). Temperature, humidity and other weather conditions during the month before the samples were taken, were associated with serum metabolites involved in sleep and depression, proteins involved in thermoregulation, and immune response genes.
“With the rich exposome and molecular information available in our catalog, we provide a valuable resource to the scientific community for finding exposure biomarkers, identifying exposure sources, improving the understanding of disease mechanisms, and, ultimately, promoting public health policies,” concludes Vrijheid.
More information:
Léa Maitre et al, Multi-omics signatures of the human early life exposome, Nature Communications (2022). DOI: 10.1038/s41467-022-34422-2. www.nature.com/articles/s41467-022-34422-2
Journal information:
Nature Communications
Source: Read Full Article