All SARS-CoV-2 variants blocked by a simple peptide with nanomolar neutralizing efficacy
In a recent study published in the journal PNAS, researchers from Finland and the United States reported a novel heptad repeat 2 (HR2) peptide that inhibited severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) without any chemical modifications.
 Study: Nanomolar inhibition of SARS-CoV-2 infection by an unmodified peptide targeting the prehairpin intermediate of the spike protein. Image Credit: Kateryna Kon / Shutterstock
Study: Nanomolar inhibition of SARS-CoV-2 infection by an unmodified peptide targeting the prehairpin intermediate of the spike protein. Image Credit: Kateryna Kon / Shutterstock
Background
SARS-CoV-2 is evolving continuously with all its variants of concerns (VOCs) harboring mutations in the receptor-binding domain (RBD) of its spike (S) glycoprotein rendering all coronavirus disease 2019 (COVID-19) vaccines and monoclonal antibody (mAb) therapies ineffective.
There is an urgent need for more effective antivirals against SARS-CoV-2, especially those targeting structures and processed less vulnerable to mutation (e.g., heptad repeat 1 and 2 (HR1-HR2) six-helix bundle of the SARS-CoV-2 S).
Previously, researchers have used several structural engineering approaches to make more effective HR2-based inhibitors based on 36 amino acid segments of SARS-CoV-2 S residues 1168–1203.
About the study
In the present study, researchers found that an unmodified SARS-CoV-2 HR2 peptide spanning S residues 1162–1203 termed longHR2_42 had much improved SARS-CoV-2 inhibiting properties relative to all previous attempts. They characterized the HR1-HR2 bundle using a molecular scaffolding method to determine its high-resolution cryo-electron microscopy (cryo-EM) structure of the longHR2_45 bound to HR1.
Next, they designed an extended HR2 peptide with a longer N-terminal region spanning amino acid residues 1159–1179 using Coot that attained SARS-CoV-2 inhibition in the nanomolar (nM) range. They refined the structure in Rosetta using the automated structure refinement protocol. Likewise, they assessed HR1-HR2 complex formation using intrinsic fluorescence on clear native-polyacrylamide gel electrophoresis (CN-PAGE).
The researchers developed two assays to detect complex formation between HR2 variants and HR1 and a cell-cell fusion assay to test the effects of the N-terminal extension of HR2 on inhibiting the membrane-fusion function of SARS-CoV-2 S.
The first assay used the cell surface of Escherichia coli to display HR2 peptides on an enhanced circularly permuted OmpX protein (eCPX)-protein scaffold. Next, the researchers incubated these cells with a green fluorescent protein (GFP)-tagged HR1 and GFP-positive cells expressing HR2 peptides that bound HR1, which they selected using fluorescence-activated cell (FAC) sorting. The second assay employed messenger ribonucleic acid (mRNA) display to detect the HR2-HR1 complex formation.
Study findings
The novel N-terminal extended HR2 peptide developed in the current study showed ~100-fold more potent inhibition of all major SARS-CoV-2 variants, with a half-maximal inhibitory concentration (IC50) value of ∼1 nM as assessed by a cell-based fusion assay. Notably, it had ∼10-fold lower inhibitory activity against the Omicron variant. Three mutations in the HR1 region of Omicron – Q954H, N969K, and L981F—all nested in the region between HR1 and the N-terminal extension of HR2, explained weaker inhibition activity of longHR2_42.
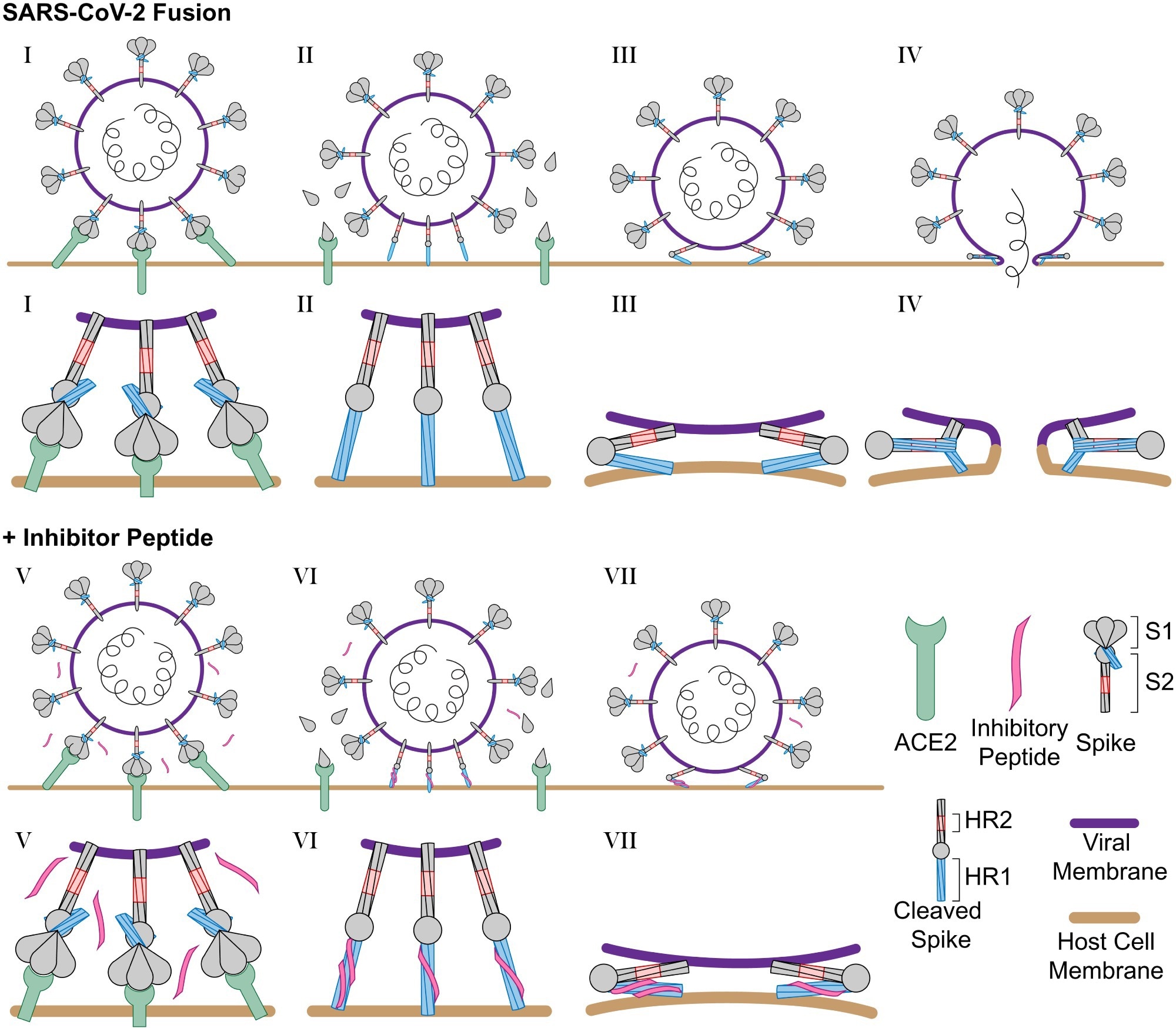 Schematic of SARS-CoV-2 infection and inhibition by HR2 peptides. (I) SARS-CoV-2 binds the host cell receptor ACE2 through an interaction with the S1 domain of the S protein. (II) After cleavage by host cell proteases, the S1 domain is released and the S2 domain of the S protein extends into the host cell membrane. (III) Triggered by slightly acidic pH, the S2 domain folds back pulling the viral and host cell membrane into close proximity, and (IV) the folding of the HR1 and HR2 domains catalyzes fusion of the viral membrane with the host cell membrane. (V) In the presence of the longHR2_42 inhibitor, the S protein engages the host cell receptor in the similar manner where (VI) after cleavage the longHR2_42 inhibitor binds the HR1 domain which (VII) prevents the folding of the HR1 and HR2 domains together and blocks membrane fusion. In this model, only a subset of the potential binding sites for the inhibitor need to be occupied in order to block fusion.
Schematic of SARS-CoV-2 infection and inhibition by HR2 peptides. (I) SARS-CoV-2 binds the host cell receptor ACE2 through an interaction with the S1 domain of the S protein. (II) After cleavage by host cell proteases, the S1 domain is released and the S2 domain of the S protein extends into the host cell membrane. (III) Triggered by slightly acidic pH, the S2 domain folds back pulling the viral and host cell membrane into close proximity, and (IV) the folding of the HR1 and HR2 domains catalyzes fusion of the viral membrane with the host cell membrane. (V) In the presence of the longHR2_42 inhibitor, the S protein engages the host cell receptor in the similar manner where (VI) after cleavage the longHR2_42 inhibitor binds the HR1 domain which (VII) prevents the folding of the HR1 and HR2 domains together and blocks membrane fusion. In this model, only a subset of the potential binding sites for the inhibitor need to be occupied in order to block fusion.
Against the Delta VOC, IC50 values varied nearly fivefold between the VSV-SARS-CoV-2 infection assay and the authentic SARS-CoV-2 infection assays.
Further, the researchers noted that longHR2_42 did not show a corresponding increase in the apparent affinity for HR1 in all three binding assays described in the study. There are two possible explanations for the same. First, these assays were not sensitive enough to detect slight changes in affinity. However, it is more likely that other factors affected the anti-SARS-CoV-2 efficacy of HR2-derived peptides.
Single virus imaging has revealed that nine to 12 S proteins are necessary for SARS-CoV-2 fusion to host cell receptors; therefore, a single inhibitory protein could prevent fusion. Also, S protein cleavage requires transmembrane protease serine 2 (TMPRSS2) or cathepsin. Since the researchers added dynamin inhibitor dynasore-OH to inhibit cathepsin in the washout experiments, it established a prehairpin intermediate of SARS-CoV-2 S for peptide binding. Thus, in the absence of TMPRSS2, the virus and peptide had a weak association, thus restricting inhibition lifetime to only 15 to 20 minutes. Nevertheless, the observed enhanced affinity of longHR2_42 was independent of host cell proteases necessary for S cleavage.
Conclusions
The HR2 helical region of SARS-CoV-2 S showed the potential to be a source of potent peptide-derived anti-SARS-CoV-2 therapeutics that would work against its VOCs and more distantly related viruses. Furthermore, the longHR2_42 peptide had a long inhibition lifetime (greater than three hours) despite washout in virus infection assays, suggesting that it targeted a pre-hairpin intermediate of the SARS-CoV-2 S protein. Indeed, the study data provided further support for the pre-hairpin intermediate of the SARS-CoV-2 S glycoprotein.
According to the authors, extending the longHR2_42 peptide sequence could enhance its potency. In addition, its sequence optimization could further improve its activity and provide a platform for developing other novel SARS-CoV-2 variant-specific peptides.
- Nanomolar inhibition of SARS-CoV-2 infection by an unmodified peptide targeting the prehairpin intermediate of the spike protein. Kailu Yang, Chuchu Wang, Alex J. B. Kreutzberger, Ravi Ojha, Suvi Kuivanen, Sergio Couoh-Cardel, Serena Muratcioglu, Timothy J. Eisen, K. Ian White, Richard G. Held, Subu Subramanian, Kendra Marcus, Richard A. Pfuetzner, Luis Esquivies, Catherine A. Doyle, John Kuriyan, Olli Vapalahti, Giuseppe Balistreri, Tom Kirchhausen, and Axel T. Brunger, PNAS 2022, DOI: https://doi.org/10.1073/pnas.2210990119, https://www.pnas.org/doi/10.1073/pnas.2210990119
Posted in: Drug Discovery & Pharmaceuticals | Medical Science News | Medical Research News | Disease/Infection News | Pharmaceutical News
Tags: ACE2, Amino Acid, Antibody, Assay, Cell, Cell Membrane, Coronavirus, Coronavirus Disease COVID-19, covid-19, Efficacy, Electron, Electron Microscopy, Electrophoresis, Fluorescence, Fluorescent Protein, Gel Electrophoresis, Glycoprotein, Helix, Imaging, Membrane, Microscopy, Monoclonal Antibody, Mutation, Omicron, Peptides, pH, Protein, Receptor, Respiratory, Ribonucleic Acid, SARS, SARS-CoV-2, Serine, Severe Acute Respiratory, Severe Acute Respiratory Syndrome, Spike Protein, Syndrome, Therapeutics, Virus

Written by
Neha Mathur
Neha is a digital marketing professional based in Gurugram, India. She has a Master’s degree from the University of Rajasthan with a specialization in Biotechnology in 2008. She has experience in pre-clinical research as part of her research project in The Department of Toxicology at the prestigious Central Drug Research Institute (CDRI), Lucknow, India. She also holds a certification in C++ programming.
Source: Read Full Article